3. SEDM Pipeline¶
3.1. Pipeline Overview¶
A new pipeline developed by Mickael Rigault has been installed for automatic reduction of SEDM data. The distribution is available on github. It generates a geometry solution from the calibration images and then automatically extracts target spectra based on the WCS solution of the guider images. Please see the documentation that accompanies the github repository. The extracted spectrum is classified using SNID and if it is a ZTF target, the ascii spectrum is uploaded to the fritz marshal. The only interactive step is to generate a final report once all the extractions have been verified.
If there is a failure of the WCS solution, or if the target is particularly difficult to model with a PSF, there are ways to re-extract the spectrum. This will be described in more detail below.
In addition to an automatic circular extraction region, we have also implemented using contours from PS1 images to define the extaction region. These are referred to as contsep extractions for contour separation. These are done in parallel to the circular (robot) extractions, but are currently not automatically uploaded to the fritz marshal. In cases where contsep extractions give an improvement over the robot extractions, they can be uploaded manually (see below).
3.1.1. Python Requirements¶
The IFU pipeline is compatible with python v2.7 and v3.6 and currently runs under the miniconda3 distribution. It requires the astroconda environment from STScI and expects the name to be ‘astroconda’:
conda create -n astroconda stsci
3.2. Automated Pipeline Operations¶
Now we describe the steps that the pipeline takes during the automated operations. We start with the pre-science processing that generates the nightly geometry solution and then continue with the science target processing.
The pipeline should be running continuously, but if minar goes down, or if
you need to restart the pipeline first cd to ~/redux and then use the
command:
~/spy ~/sedmpy/drpifu/AutoReduce.py --wait |& tee -a run.log
The --wait option is used when you re-start the pipeline before any new
data are taken. If you need to re-start the pipeline after observations have
begun, then leave out the --wait option. There are two aliases that
perform these functions:
autored
and
autored_wait
3.2.1. Pre-Science Processing¶
Pre-science processing occurs in the afternoon and takes roughly 30 minutes to complete. In the afternoon when the UT date changes, the following steps are automatically performed:
- The appropriate reduced directory is created using the UT date:
/scr2/sedmdrp/redux/20230407(e.g.)
The required raw calibration files are linked into the directory as they are taken.
Once all the bias files are acquired the master biases are generated.
All subsequent calibration files are linked into the directory and then bias-subtracted and cosmic ray cleaned.
Once all the calibration files are acquired a geometry solution and flat field is generated.
If there is a failure in the geometry solution, geometry files from previous runs are linked in.
The most recent fluxcal fits file is linked in, usually from the previous night.
You can verify the success of the geometry solution and flat field by looking at the plots generated when they are produced. One is the wavelength solution dispersion and the other is the flat field. They will be in the reduced directory and have names that start with the UT date string (20230407, e.g.).
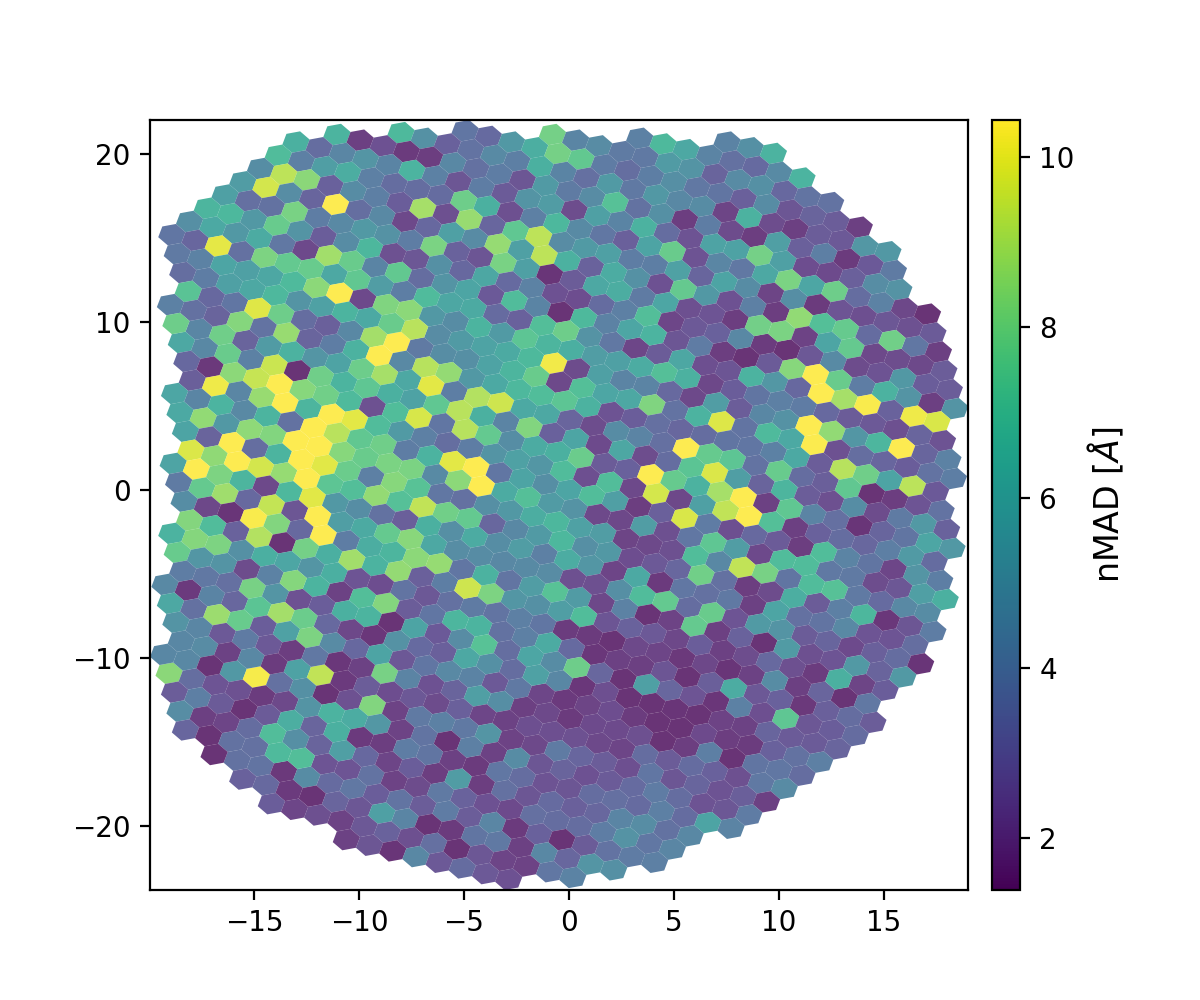
Figure 1. Wavelength solution dispersion in Angstroms (20230407_wavesolution_dispersionmap.png).¶
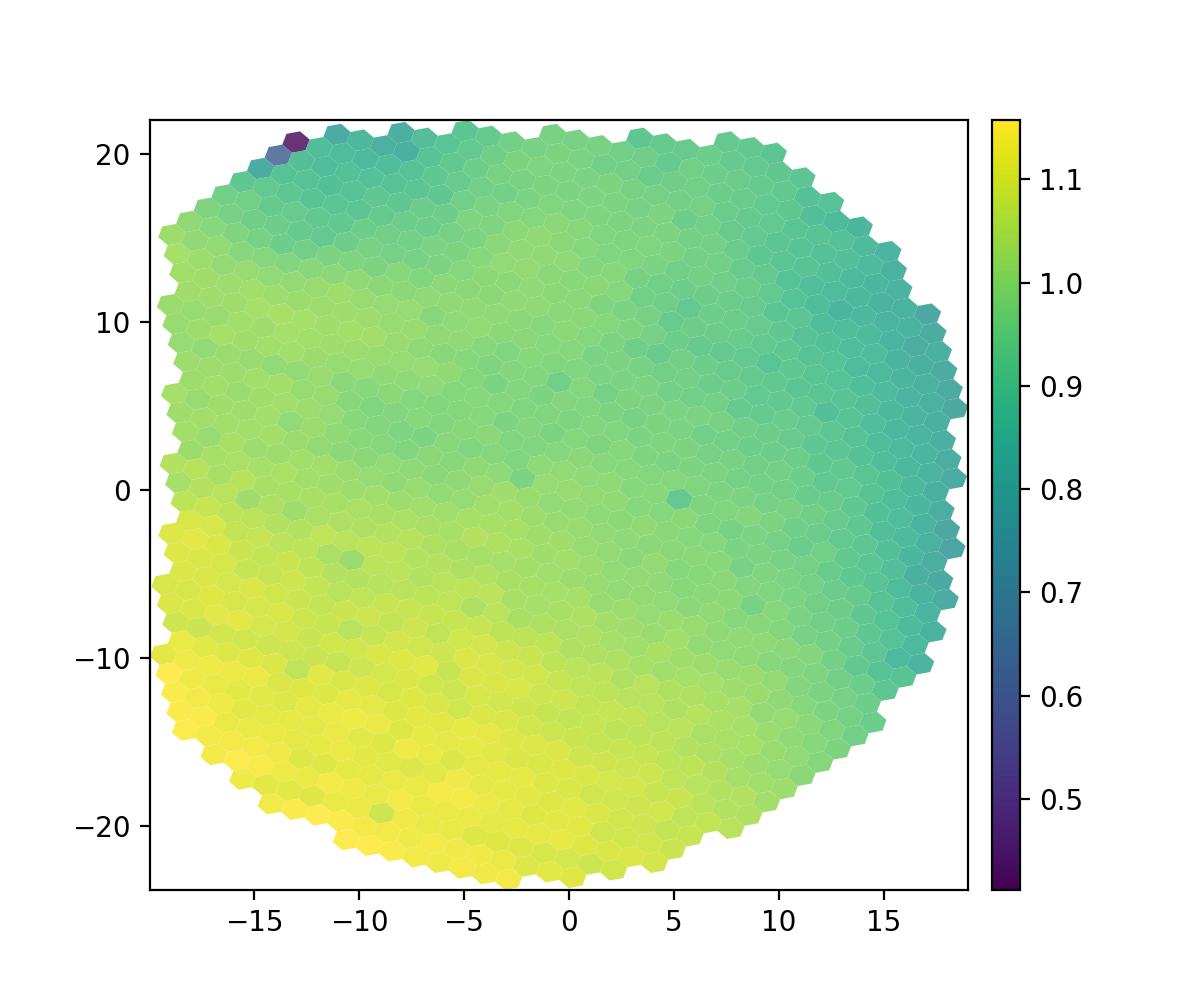
Figure 2. Flat field (20230407_flat3d.png).¶
3.2.2. Science Processing¶
Now the SEDM is ready for science images. Near the end of astronomical twilight, science image acquisition begins. The following steps are automatically performed:
All new IFU images are linked in and bias-subtracted and cosmic ray cleaned.
The sky traces are used to solve for the flexure offsets for the observation.
The geometry solution is used to generate a flexure-corrected cube for the observation.
- If the target is a standard star:
an astrometry image is generated.
the brightest spaxel is used to define the extraction region.
PSF-forced spectro-photometry is performed.
a fluxcal fits file is generated.
- If the target is a science object:
an astrometry image is generated from all the guider frames and the WCS is solved.
an extraction region in the IFU is based on the guider WCS and the target coordinates.
a separate extraction using the contsep method is created.
PSF-forced spectro-photometry is performed.
the most recent fluxcal file is used to calibrate the science target.
the telluric absorption is corrected based on header AIRMASS.
the resulting spectra are classified using SNID
the SNID results are put in the ascii and fits spectrum headers.
the extractions are recorded in a file called
report.txt
- If the target is a ZTF object:
the robot spectrum is uploaded to the fritz marshal.
the marshal URL is recorded in the file
report_ztf_fritz.txt
The format of the ascii spectrum that is generated is universal enough to be input to any classifier (Superfit, e.g.).
3.2.3. Quality¶
A quality value is assigned to each spectrum based on several criteria. In this scheme, quality values 0, 1, and 2 are considered acceptable. Any value above 2 indicates a major problem and the spectrum will neither be classified nor uploaded to the marshal. Here is the current quality scheme:
Quality 0: default quality
Quality 1: extraction OK (usually recovered from Quality 5)
Quality 2: a minor problem was encountered (currently not used)
Quality 3: the telescope offset placed the target outside the IFU
Quality 4: more than 20% of the flux is negative
Quality 5: the guide image astrometry failed (for science targets only)
Standard star observations always use the brightest spaxel to define the centroid. Therefore, standard stars should never have a quality of 5.
Quality 3 objects cannot be fixed. Quality 4 and 5 objects may possibly be fixed, but will require hand-extraction (see Adjustment below).
3.2.4. Standard Star Verification¶
The first on-sky IFU observation should be a standard star, usually taken during the end of nautical twilight. The resulting calibration will be used for the rest of the night, therefore it is important to verify that it is a successful observation. This new calibration will replace the one that was linked in by the pipeline prior to opening the dome. Only a quality 0 standard star observation will be subsequently used, so if a bad observation results in a quality 4 observation (the most likely), the pipeline will simply use the calibration that was linked in before and this will be fine.
If, for some reason, this new calibration is bad, but the quality is still 0, it will mis-calibrate the entire rest of the night, resulting in a lot of work to re-extract and re-calibrate all of the night’s observations. Thus, it is worth checking the quality of the first standard observation as it happens.
3.2.4.1. How to Handle a Bad Quality Zero Observation¶
If the first observed standard looks bad, but has a quality of 0, there is a simple procedure for removing the bad calibration so it will not be used. Here is an example of such an observation:
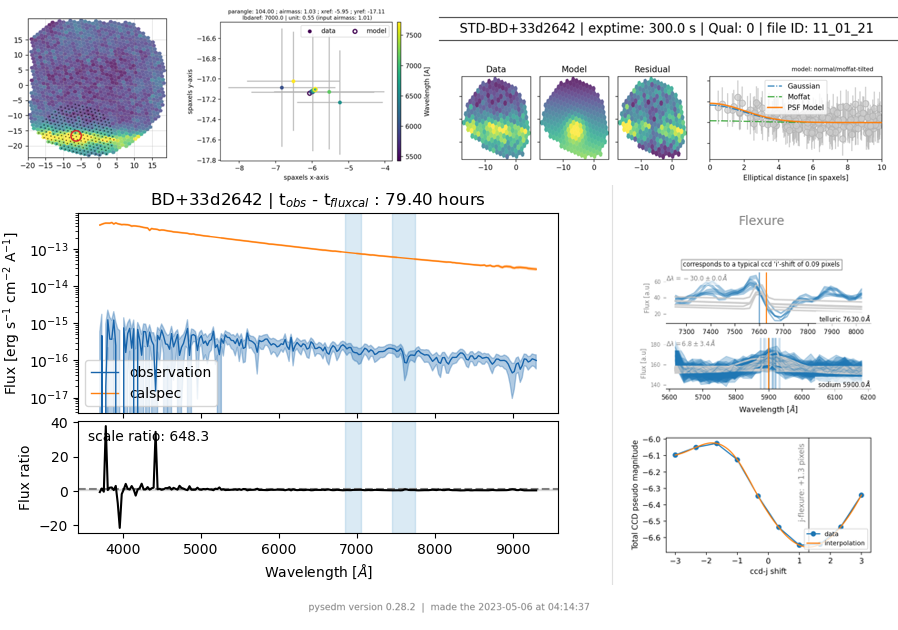
There was a tracking problem, but the pipeline thinks it was extracted correctly, even though it has spikes in the spectrum that will mis-calibrate the science observations.
See the next section for how to log into the DRP machine. This is where you will fix the problem. You can either ssh into the machine or use the VNC. You should have enough time before the first IFU science image will require the calibration file.
Essentially, what you do is just move the bad calibration aside as follows:
> cd ~/redux/20230506
> mkdir bad
> mv fluxcal_auto_robot_lstep1__crr_b_ifu20230506_*STD-*.fits bad
Be sure to substitute the correct day number for the day numbers in this
example. The pipeline will not find this calibration in the bad directory
and the linked-in calibration will be used instead.
3.3. Interactive Processing¶
All target extractions should be verified and adjusted if required. Once that is done a final report is generated that sends out a summary e-mail of the night’s results. In order to do this, one has to connect to minar.caltech.edu via a VNC connection. If the screen lock is active, just enter the password to unlock it. Below is is a figure showing the layout of the main desktop screen connected through the VNC connection.
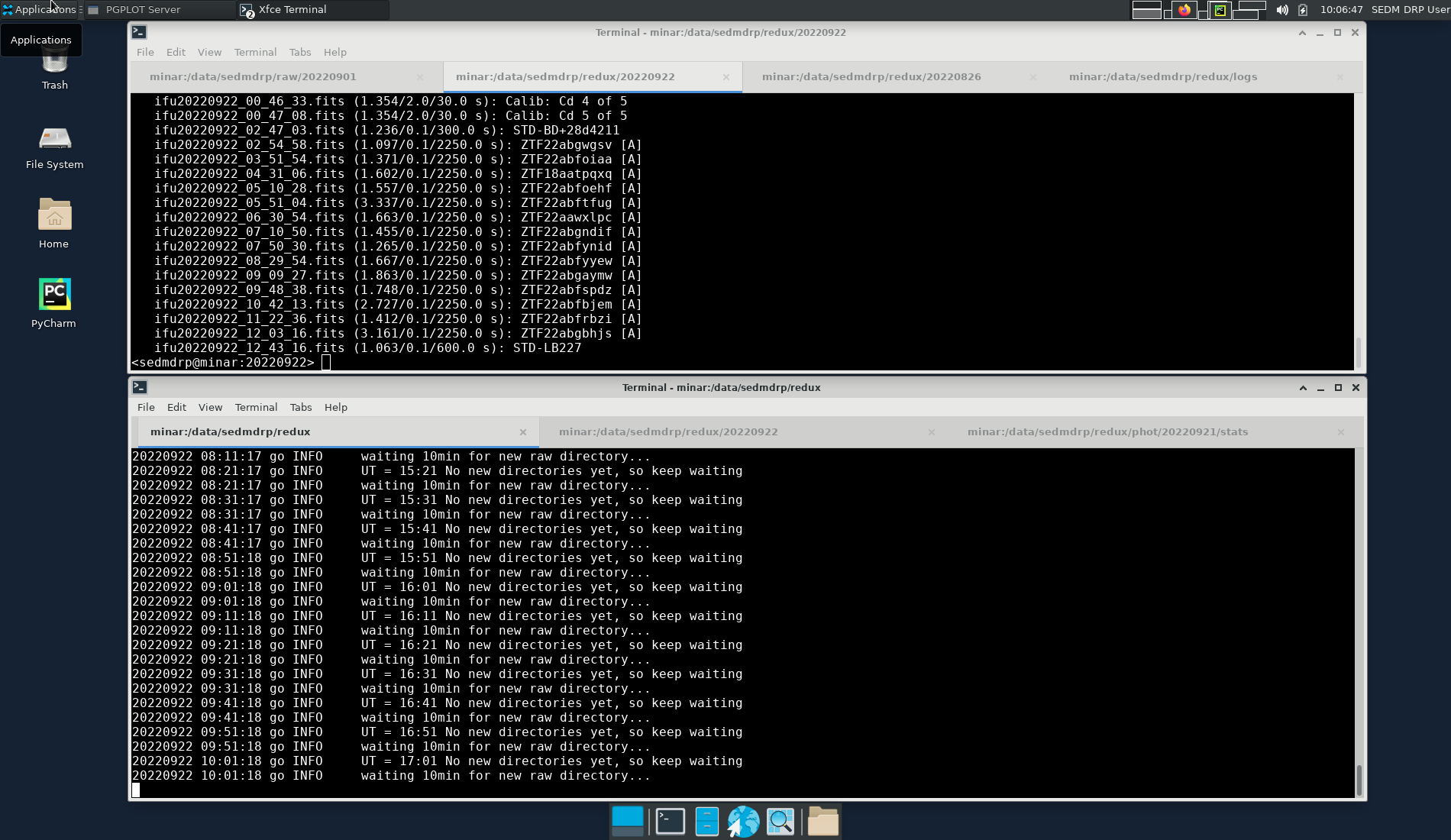
Figure 3. minar sedmdrp desktop on screen 7 (5907).¶
The automatic pipeline script is running in the bottom xterm window in the leftmost tab. Some status information can be gleaned from the output there. The second tab on the top xterm can be used to examine the what.list file. The middle tab on the bottom xterm may be used by the observer to examine the files on minar. A web browser will be set up on the secondary desktop to the right which can be selected using the chooser on the upper right. This is where you can interact with the SEDM web site and the fritz marshal and other web services to look at finder charts.
In the middle tab of the bottom Xterm window, the observer interacts with the pipeline as described below. Be sure to cd into the current directory, which is the UT date formatted as YYYYMMDD (20230407, e.g., which would be found in /scr2/sedmdrp/redux/20230407).
3.3.1. Verification¶
The automated pipeline generates verification plots as each image is processed.
These are PNG image files that start with verify_. You can display all
of them using the display command from ImageMagick like this:
display verify_auto_robot_*.png &
Figures 4 - 6 show the three types of verification plots. For all three types, the acquisition finder chart is shown in the upper right and the IFU spaxel plot is in the upper left. The PSF extraction results are shown in the lower left in three plots showing the Data, Model, and Residual. Finally, in the lower right, is shown some form of the extracted spectrum. For a standard star, it will show the calibration check plot comparing the reference spectrum to the observed spectrum (see Figure 4).
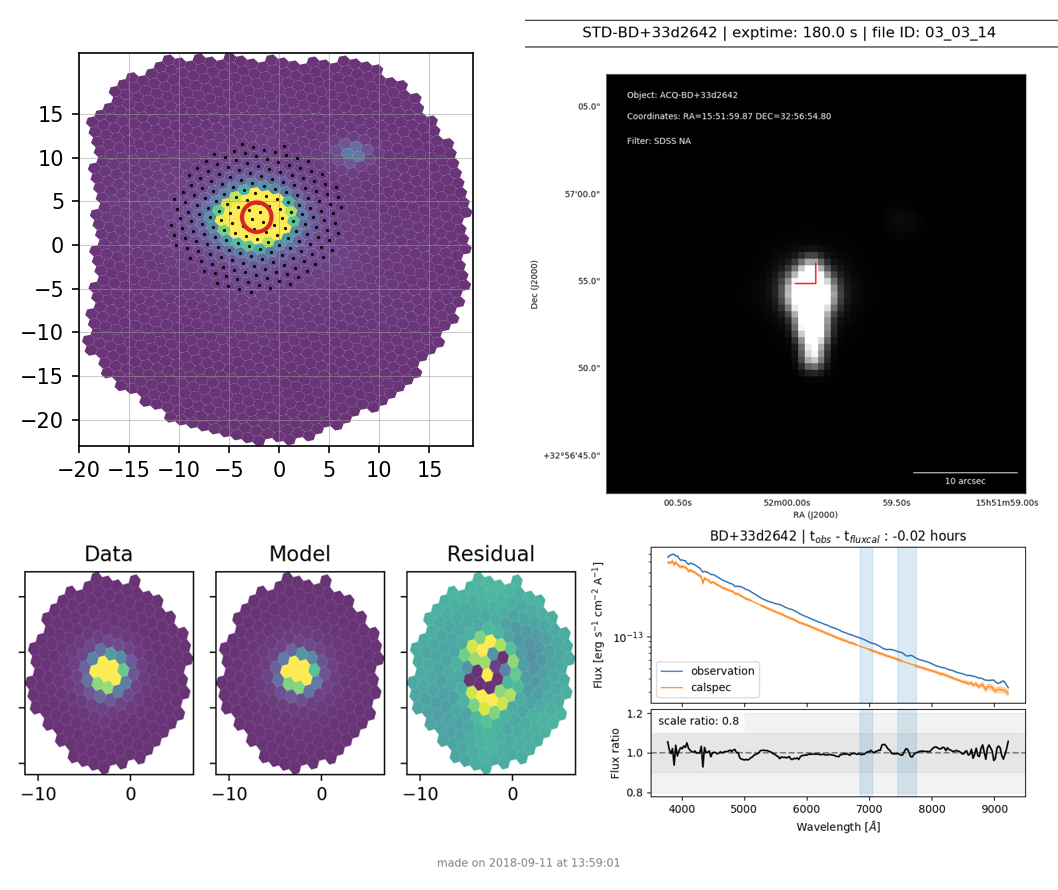
Figure 4. Verification plot for standard star BD+33d2642¶
For a science target that has a successful classification from SNID, it will show the SNID template match plot (see Figure 5).
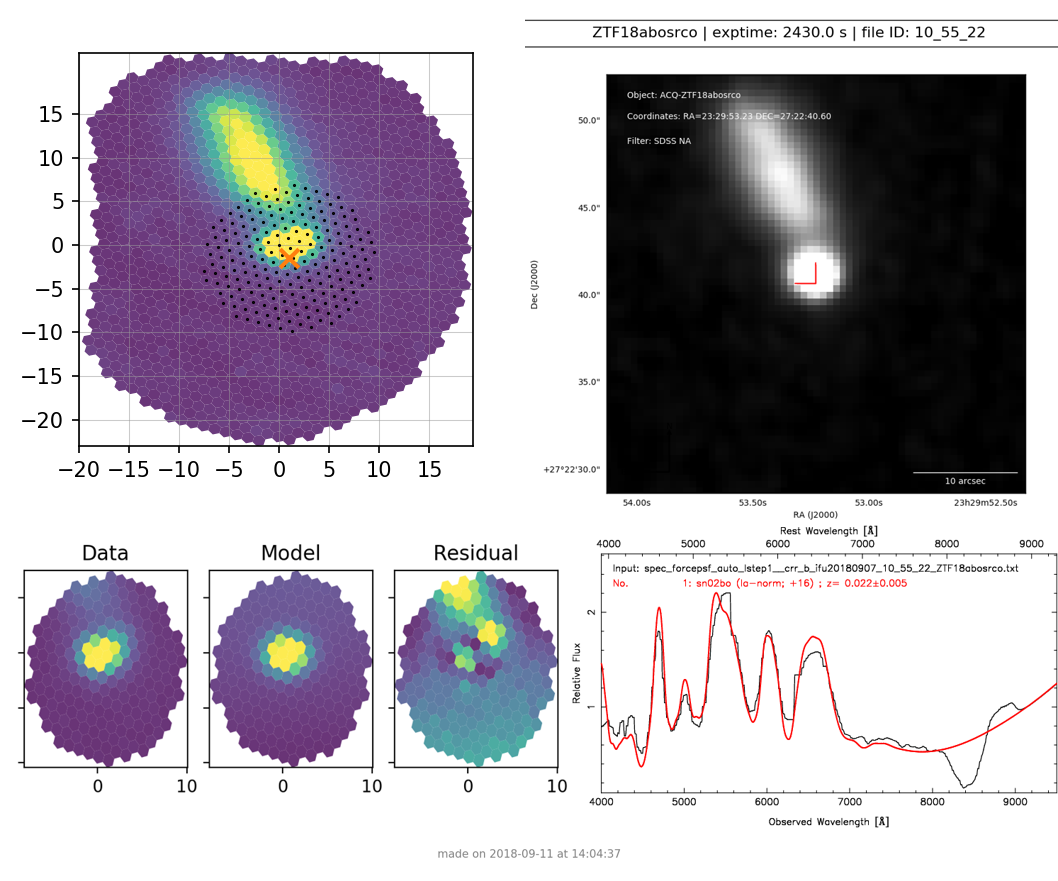
Figure 5. Verification plot for successfully typed science target ZTF18abosrco¶
For a science target for which SNID fails to find a classification, it will show only the extracted spectrum (see Figure 6).
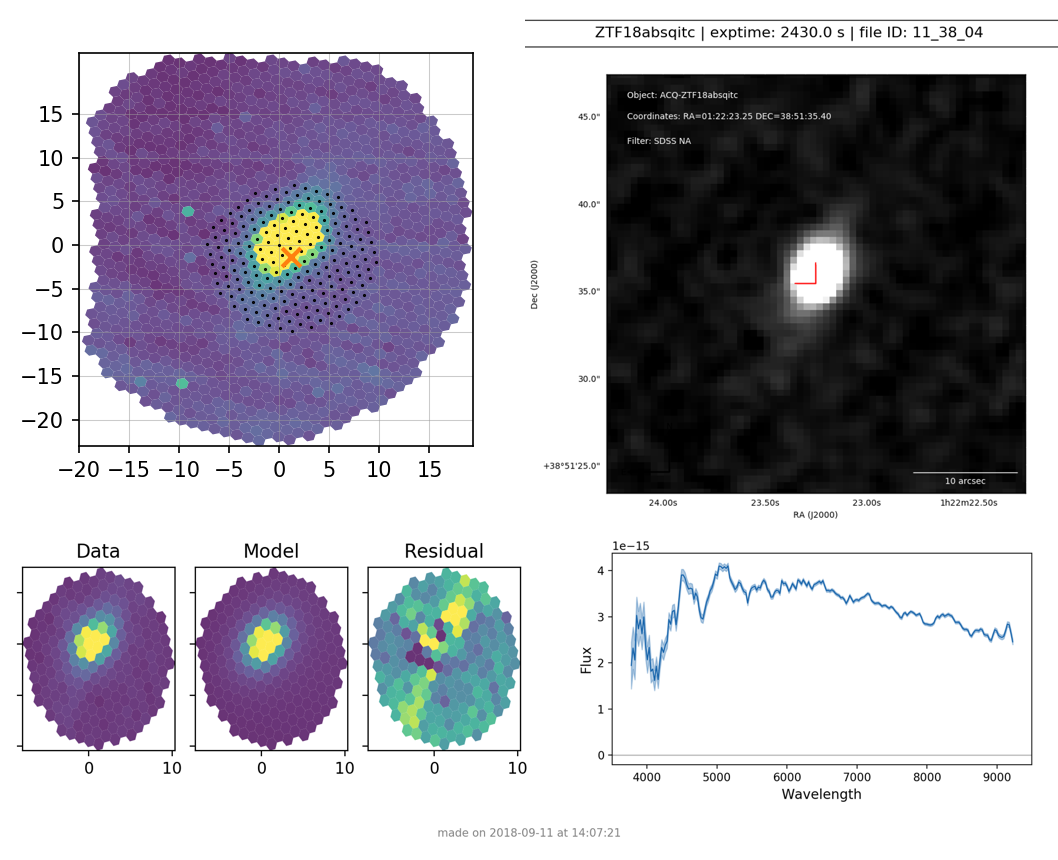
Figure 6. Verification plot for unsuccessfuly typed science target ZTF18absqitc¶
Now that we are also using contsep for extractions, you will want to display those verification images separately as follows.
display verify_auto_contsep_*.png &
These look the same as the previously described verification images, except for the contsep in the file names and the contours used for extraction will sometimes look different.
The first step of verification is to compare the B&W finder (upper right) with the IFU extraction region (upper left). The red right-angle in the B&W finder indicates the location of the target. If the IFU extraction region indicated by black dots contains the object and the centroid, indicated by either a red X or a red circle is reasonably close to the target, then this is probably a good extraction. Next, examine the PSF fit and residual plots in the lower left. If the model looks reasonably close to the data and the residuals look like the model accounted for most of the target’s flux, then the extraction was successful. This is also bolstered if the spectrum looks good and is either a good match to a SNID template, or to a reference spectrum, or seems to have good signal-to-noise.
If you want further verification of the target, you will need to move to the desktop to the right (using the chooser in the upper right). There you can open a web browser, if needed, and log into the ZTF fritz marshal, the TNS website, or any other web-based source of finder charts for the target.
3.3.2. Compare Extractions¶
You will want to also compare the robot and contsep extractions to see if the contsep extraction provides an improvement in host subtraction over the robot extraction. Check the spaxels used to define the extraction region. If the contsep spaxels (indicated with the black dots) do a good job of excluding host spaxels, then you will want to upload the contsep spectrum to the marshal. In many cases, the extractions of the robot and contsep methods are the same, especially if either the target is right on top of the host nucleus, or if the target is well separated from the host. In these cases, you don’t need to do anything. In the case where the contsep is clearly better than the robot extraction, then upload the contsep spectrum to the marshal. Please follow this example, substituting the correct contsep spectrum file.
fritz spec_auto_contsep_lstep1__crr_b_ifu20200710_04_14_53_ZTF20abjnmhn.txt
This command will upload the contsep extracted spectrum to the marshal.
If the contsep extraction is no better than the robot extraction and you still feel the robot extraction can be improved, follow the instructions in the next section.
3.3.3. Adjustment¶
There are a few types of adjustment that can be made, depending on the particular situation. We will describe the most common ones below.
NOTE: if the target was given the quality value of 5 (guide image astrometry failed), then you must identify the target by hand and reset the centroid appropriately. See the section below on how to recover a quality 5 spectrum in which the intended target was extracted correctly.
3.3.3.1. Redex Script¶
There is a script available that performs many of the bookkeeping tasks
required by re-extraction. It is called redex and can be used as follows:
redex <timestr> [<X Y>],
where <timestr> is the UT time stamp for the specific observation, and <X Y> are replaced by the corrected centroid values as determined from the IFU spaxel plot. The <timestr> is formatted:
HH_MM_SS,
and is shown in the title of the verification plot. The <X Y> values are
optional, and if not included will invoke the --display option for
extract_star.py. The script does the following:
generates a redo timestamp tag based on the current local time:
redoHHMMSS,prompt for the user’s name (defaults to env var SEDM_USER value),
re-run extract_star.py with the appropriate parameters,
regenerate the spectrum file for the object with the redo timestamp tag,
re-generate the extraction plots with the redo timestamp tag,
remove any old classification files generated by SNID,
run SNID on the new spectrum,
generate and display a new verification plot with redo timestamp tag (type
qto proceed),- prompt user to either accept or reject re-extraction,
if rejected, delete all files with redo timestamp tag and exit script,
if accepted continue with items below,
generate new pysedm_report plot,
- push this plot to the SEDM-P60 slack channel pysedm-report,
(if you add
--localto command line, this won’t happen),
- if it is a ZTF object, upload new spectrum to the marshal,
(if you add
--localto command line, this won’t happen),
- update minar DB tables,
(if you add
--localto command line, this won’t happen).
3.3.3.2. Recover a Quality 5 Spectrum¶
Sometimes, even if the astrometry fails, the target will be the brightest object
in the IFU. In these cases, the extraction will be correct, but it will have a
Quality of 5. To fix this, just add --recover to the call to the redex
script:
redex <timestr> --recover
This will update the value of Quality for the extraction to 1 in the spectrum files and the minar DB, and will upload the updated spectrum to the marshal, if it is a ZTF object. Since you have already determined that the extraction is correct, no plot is displayed and you will not be prompted to approve it.
This command now has an alias on minar in the sedmdrp account:
recover <timestr>
3.3.3.3. Adjust Centroid¶
This is the simplest adjustment to make. It will arise in some cases if the WCS solution of the guider images failed (Quality 5). This is indicated in the IFU spaxel plot when the centroid is plotted a red circle instead of a red X. When the WCS solution fails, the extraction is defined by the brightest pixel. This is fine for standard stars, but does not always work for science targets. Sometimes even successful WCS solutions will define the centroid in the wrong place. Let the finder chart in the verification plot and any other finders from the web be your guide.
It is also possible that a target that is strongly influenced by a neighbor (host galaxy, nearby star) can be fixed by just moving the centroid, and hence moving the extraction region, off of the offending neighbor.
To make this adjustment, you simply need to pass the new centroid to the redex script. Use the IFU spaxel plot to determine the new centroid for the target. Then enter the command:
redex <timestr> <X Y>,
using the parameters described above. Here is an example:
redex 10_55_22 0 -5.
The script will display the new verification plot that will allow you to assess if your new position had the intended effect. This plot will now have a black cross where your adjusted centroid falls on the spaxels.
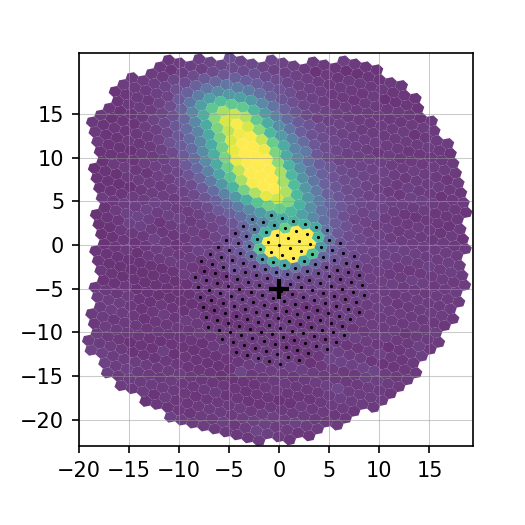
Figure 7. Adjusted centroid indicated by black cross.¶
It is fine to tweak the centroid and re-extract the spectrum more than once. It’s important to get a good extraction and this sometimes takes more than one adjustment to the centroid. Just be sure to reject the extraction until you get an extraction that looks good.
NOTE: passing the centroid to the redex script will remove the quality 5 condition.
NOTE: there is nothing in the verification plot for this object to indicate that it needs adjustment. This was done just to demonstrate the procedure.
3.3.3.4. Adjust Extraction Region¶
This is also a fairly easy adjustment to make. If the extraction region includes a neighbor that strongly influences the psf model, and just moving the centroid doesn’t fix it, you can use the redex script to invoke the –display parameter of the extract_star.py program to re-draw the region. To do this enter the command without centroid values:
redex <timestr>,
which will bring up a display window showing the IFU spaxel plot with the region and the right is the spaxel map where you can re-draw the region.
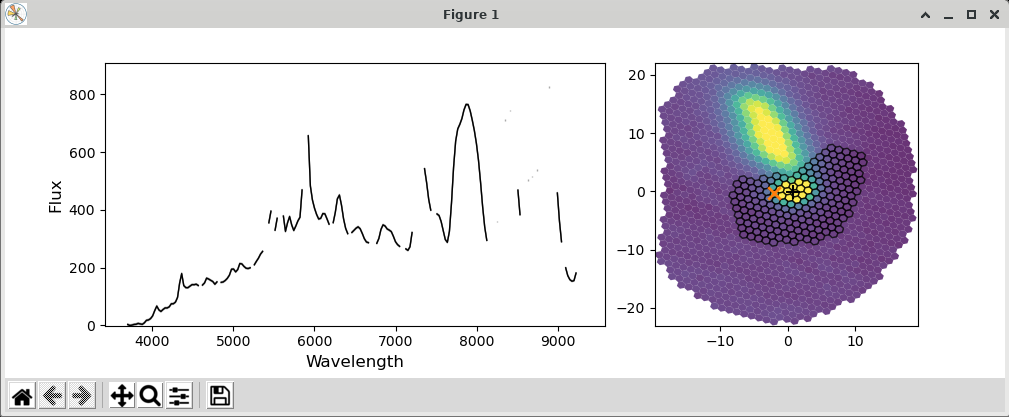
Figure 8. extract_star.py with the --display parameter and a hand-drawn extraction region.¶
Just hit the shift key and draw a region (by left clicking and dragging the mouse) around your target that does not include the offending neighbor. Once you release the left mouse button, the selected region will be shown on the plot (see Figure 8). If you want to try again, hit the <ESC> key, which will reset the region, and try again. If you want to use a new centroid, just double-click on the location of the new centroid. This will be required, if the target was assigned a quality of 5 (guider image astrometry failed). Once you are happy with the centroid and region, close the plot. This is done by clicking the ‘X’ in the upper right corner of the display window. The extraction will proceed once the window is closed.
Here is the command that produced Figure 8:
redex 10_55_22.
The script will display the new verification plot so you can either accept or reject this re-extraction.
NOTE: if the target was assigned a quality of 5, you will have to double-click on the target to reset the centroid. If you do not, the target will still have a quality of 5 and won’t be classified or uploaded.
3.3.3.4.1. Fix A Cosmic Ray¶
Using the --display parameter also allows you to find and avoid spaxels
that are corrupted by a cosmic ray. After the redex command is entered
(without centroid values), you can click on individual spaxels until you see
the one that is heavily influenced by the cosmic ray. Then, hit the shift key
and draw your extraction region so as to exclude the offending spaxel. You may
have to expand the window to more accurately draw the region.
3.3.3.5. Use Coarser Sampling¶
The extract_star.py script called by the redex script samples the
wavelengths in binned steps specified by the --lstep parameter. The
default value is one, but if the noise level is high, one may try a larger
binning by specifying it on the redex command line. For example:
redex 10_55_22 --lstep 2
will sample the wavelengths at twice the bin size as the default. One has to exercise caution when doing this because narrow emission lines can be strongly impacted.
3.3.3.6. Adjust Extraction Method¶
This is a more challenging adjustment to make. As of now, the two previous adjustments seem to be able to fix nearly every situation. If you need to perform an aperture extraction, please contact the SEDM team and we can instruct you how to do this.
3.3.4. Final Report¶
The last step at the end of the night is to generate the final report which sends a night summary e-mail report out the to the SEDM team. To initiate this final step, please enter:
make report
make finalreport
This last command will now prompt you for a comment about the night. Refer to the seeing monitor or the night statistics page on the minar website and briefly record the quality of the seeing and conditions for the night in your comment. After the report is sent out, the script will do some cleaning and gzipping of files to conserve disk space. Remember that if you want to re-extract an object after this step, you will have to gunzip the appropriate crr_b_ifu*.fits.gz file.
It is a good idea to check the e-mail (if you are on the list) and make sure all of the links work and that the correct extractions are displayed.
Congratulations! You are done, for now…
Last updated on 17 May 2023